Recently, PatientsLikeMe sent a message to our members about an opportunity to participate in the Impact of Personal Genomics (PGen) Study, led by researchers at Harvard and the University of Michigan. Each of the 1,000 study participants receives personal genomic testing services at a significantly subsidized discount. Using a series of surveys, the investigators will then look at the risks and benefits of learning this information. What, for example, will participants do with their discoveries? Will they make health behavior changes? Will they tell anyone – and if so, who?
PatientsLikeMe member PF Anderson decided to not only join the study, which has now reached its maximum enrollment, but to also start a blog about her experience. Find out why and much more in our interview below.
1. What led you to participate in the PGen study?
The “why,” for me, had three main drivers. First, I’ve been ill for over a decade, and only recently tracked it down to what appears to be celiac disease, but all the blood tests have come back negative for both celiac and gluten-intolerance or wheat allergy. Second, I am both a medical librarian and an emerging technologies librarian. I firmly believe in supporting research both by doing it and also by serving as a subject when appropriate. This project matches my professional interests in several areas, not the least of which has been an emerging awareness of the essential nature of personal genomics and big data to moving research and discovery forward, especially in the areas of rare diseases. Third, curiosity. My family seems to be packed to the gills with various genetic conditions, and I’ve always thought we’d make an interesting study!
2. Talk about the decision to blog about the study as well.
Why blog? Because, assuming that this IS an important area for research to change the lives of real people, then it is absolutely critical to educate, inform, and openly dialogue about the risks and benefits of personal genomics. Some people I talk with are quite worried about some aspects, while others aren’t even thinking about potential risks! I’m in the middle. I know there are risks, I’m aware that the benefits we hope for in the long term aren’t here yet, but I believe we have to start somewhere to get where I hope we’re going, and someone has to take the risks to hopefully shift the balance.
I hope that the more of us do this, and talk about it, that this will help other patients and individuals think through why they would or would not want to learn this. Also, while I am enormously impressed with the WeConsent.Us website for its information about the risks and benefits of personal genomics and sharing personal data, there is something about telling a story from a real person in their own words that has more impact. Hopefully my own thoughts and story will enrich what WeConsent.Us is doing.
3. You recently received your genetic results. What’s that been like?
Frankly, the results so far have been pretty disappointing. There are two branches of the study, one using the testing service Pathway Genomics, and the other using the testing service 23andMe. Each of the two companies runs saliva scans for different conditions, as well as other information. The primary conditions of interest for me were not included in the scan by Pathway Genomics. The results were interesting, but not very relevant. What was most interesting is that, according to the results, for the flock of conditions that run in my family, I am not at risk for ANY of them, including some I am already diagnosed as having. That seems rather surprising, so I am puzzling this over. I suspect that these are actually related to the core condition [celiac disease] that the scan didn’t include.
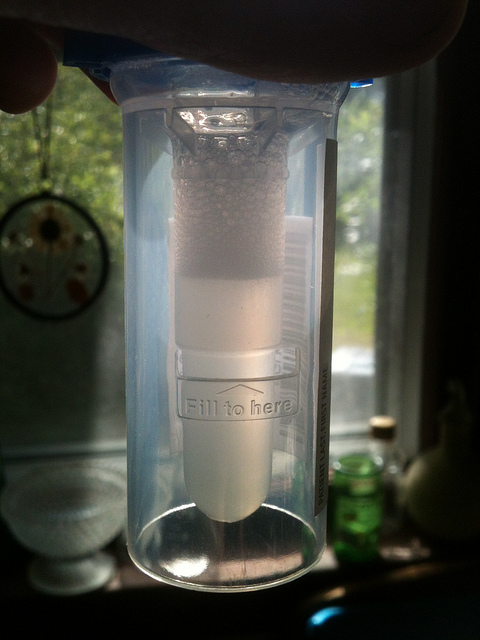

I am also a little worried that if I share my test results with my doctors, and the results show that I “don’t have” or am not at risk for these various conditions that run in my family, that the healthcare team might be less vigilant in monitoring these. Because of worrying about the risks of my healthcare team misunderstanding the results and needing the celiac test, I decided to actually spend my own money on getting the other [23andMe] scan. I’m nervous about the money, but I really feel that the information from the one test is incomplete without the other, and the risks of the incomplete information undermine the value of the original test.
4. At PatientsLikeMe, you’ve been able to chat with other PGen study participants in the forum. What have you gained from that?
The forum conversations have been fascinating! It is really interesting seeing what sort of questions other people have, their reasons, their assumptions, the information that they bring to the table. Many of the conversations there have triggered new questions for me, and opportunities to learn more.
5. If you had to pick one key takeaway from participating, what would it be?
We aren’t “there” yet, but if we ever want to get “there,” we need to start somewhere, and that’s here and now.

Pingback: Interviewed at Patients Like Me | PGen Participant
I also submitted my sample to Pathway Genomics and came to the same conclusion as PF did: “according to the results, for the flock of conditions that run in my family, I am not at risk for ANY of them, including some I am already diagnosed as having.”
I think this is for two reasons:
1. The test was a very simplistic test – ie of SNP’s (Single Nucleotide Polymorphism) (“a DNA sequence variation occurring when a single nucleotide — A, T, C or G — in the genome differs between members of a biological species or paired chromosomes in an individual” quoted from Wikipedia). In other words, they only tested one nucleotide at a time out of the many that make up a gene, which ignores all the instances in which variations in multiple nucleotides in a gene correlate with a specific disease rather than just one.
2. Initial results of the ENCODE project were just published for the first time (Sept. 2012) that suggests that the large portion of the genome outside of encodable genes is not a waste land, but in fact made up of millions of elements that control the behavior of the genes that we can now map and study, which might explain how two persons can have the same gene structure(s), including the same variations, and one affected by the sometimes related disease but the other not so affected.
In my opinion (having never taken a class in Biology), other than for a very few diseases, any attempt to diagnose diseases based on a study of SNP’s is just a waste of time and money because the relationship between the genome and a disease is far more complex than that.
With all due respect to PatientsLikeMe, I was disappointed that PatientsLikeMe would participate in such a study that misleads volunteers into thinking, like PF Anderson did and like I did, that some personally useful information could be expected to come out of it.
Dear Dave,
Thank you for your comments, and thanks to PF for sharing your experience. I appreciate your accurate and succinct assessment of the value of SNP genetics, especially for those who are already dealing with disease. This is a topic I am passionate about and have been involved in for some time. It comes down to how predictive is your ancestry and, as full genomics comes on line, how predictive are your individual genes for your individual outcomes.
I believe that for most of us they are neither very predictive nor very useful. I do, however, think they are valuable tools for looking at the common genetic basis of some diseases. They may also help to discover protective variants in the gene pool that could inform the development of drugs.
PatientsLikeMe previously gave each of its employees a full 23andMe genomics profile so that we can learn firsthand about the experience, and about the impact of genomics and biomarkers on our understanding of disease as individuals and groups. This specific study tested the method of working directly with patients to learn about their genetics. It represents a new way of doing research that 23andMe has pioneered. Now, real advances may come from these direct-to-patient approaches.
We fundamentally believe that we need to work collaboratively with patients in the research process. That means having dialogs like this, and getting feedback on where we succeeded and failed. We try to be very careful to articulate that most research is forward looking, and that we hope to learn something that will help us understand disease vs. something that would help the individual. If we failed to be clear about this, I do apologize, and will look to ensure there is no confusion in the future.
In the end understanding disease is about measuring, and measuring quickly. We all want to find predictive markers like viral load in HIV that can quickly indicate an individual on a treatment is improving or getting worse. If we had those markers in MS, ALS or Fibromyalgia, or even in mood disorders, it would dramatically improve our ability to diagnose, manage and develop treatments. Our read is that these technologies will come not from measuring DNA, but rather from measuring the state of the patients and their systems like RNA, Protein, or even gut bacteria. This experiment is the first among many designed to evaluate how well PatientsLikeMe and patients can collaborate with these technologies to validate that they actually do help understand and treat disease.
In this case I agree it’s unlikely that a few selected snips are going to affect how you manage your conditions, except perhaps in cases where genes control drug responses. The fact is we are learning more as we collaborate with patients about how to research with them rather than on them. If we work out the kinks, we can accelerate discovery, lower costs and, in the end, help make new treatments and new ways of managing disease all the more available. I think we can all agree that these are very important goals.
Jamie
PS: If you want to learn a little more about our take on personal genomics you can watch a talk I gave at MIT on this topic on this blog post here
Just a thought about the genetic links with psoriasis. My husband has a whole family who have a variety of conditions, mostly related to psoriasis. Having carried out loads of reading and research there seems to be a huge link between the conditions of psoriasis, Chrone’s Disease, MS, digestive problems, psoriatic arthritis and lack of the ability to absorb vitamin d.
Apparently there is a genetic marker that seems to connect all these conditions. We’re still hoping for more research that shows there is a loink and eventual treatment.
As a researcher, I am happy to see volunteers as passionate as PF Anderson. It is a positive attitude to participate in a research project and understand that we are not “there” yet but we will get “there”. These kind of participants are golden. I think it is because of PF Anderson’s expertise in his field. Everyone should learn about his/her personal genetics. However, they have to learn about it with the correct attitude. Education on how participants should interpret the results is also extremely important.